The fastest way to understand a VCF before reading a single variant.
Structured header parsing and standalone HTML reporting for daily review of pipeline outputs.

VCF headers contain the metadata needed to understand a file before any variant-level interpretation begins. In raw form, that metadata is difficult to inspect quickly, validate consistently, and share with others. Reference assembly, field definitions, filters, samples, and caller-specific tags are present, but not arranged for routine review.
VCFheader does not analyse variants.
It does not replace downstream interpretation tools.
It reads the header only.
It structures the metadata and writes a portable HTML report.
VCF or VCF.gz file → header parse → structured summary and HTML report → faster review, validation, and sharing
Downloads
| OS | File | SHA256 |
|---|---|---|
| Linux x86_64 | vcfheader_v0.1.0_linux_x86_64.tar.gz | 01f5cd3c3f28fce55b9726e9ed0bc0739e84182a2457adc204bb7e985fe41e93 |
| macOS universal | vcfheader_v0.1.0_macos_universal.tar.gz | 81d82e0d90716a6bdb8237073ad55be00501b6db74935f9f6029d303bcfe4c57 |
| macOS x86_64 | vcfheader_v0.1.0_macos_x86_64.tar.gz | e51e90a3eda29f9c4daa85b8d7cc83323bf708c23a07382b3be3bd50f39c529b |
We recommend the macOS universal binary for all Apple users. It runs natively on both Intel x86_64 and Apple silicon arm64 systems.
R package
Source: cran.r-project vcfheader (for installation see below).
Test VCF file
Click here to download a small test VCF file: simple.vcf (Source samtools hts-lib)
When VCFheader is most useful
VCFheader is most useful when a new file arrives and the first task is orientation. It helps users confirm the reference genome, inspect INFO and FORMAT definitions, review filters, check sample content, and identify caller-specific metadata before downstream analysis begins.
It is also useful when outputs need to be tracked across runs, tools, or pipeline versions. A standalone HTML report makes header context easier to review, archive, and share with collaborators.
Command line installation
VCFheader is distributed as a standalone binary and as an R package. The binary runs directly from the extracted directory and requires no system installation or administrator privileges. It is compatible with macOS, Linux, HPC environments, and shared compute clusters.
macOS and Linux binary installation
Download, verify checksums, and extract:
shasum -a 256 vcfheader_v*.tar.gz
tar -xzf vcfheader_v*.tar.gz
cd vcfheader
./vcfheader --help
(See note below for macOS warning)
Run directly:
./vcfheader report simple.vcf --out simple_vcfheader.html
macOS note
We provide the universal macOS binary runs natively on both Intel x86_64 and Apple silicon arm64 systems.
Downloaded binaries may be quarantined by macOS. Remove the quarantine flag if prompted:
xattr -d com.apple.quarantine vcfheader
Linux and HPC note
VCFheader can be used from any directory with execute permissions. Administrators may optionally expose it via an environment module:
module load vcfheader
Quick start for command line
Generate an HTML report from a compressed VCF:
./vcfheader file.vcf.gz --out file_vcfheader.html
Generate an HTML report from an uncompressed VCF:
./vcfheader file.vcf --out file_vcfheader.html
Minimal run using default output naming:
./vcfheader file.vcf.gz
VCFheader reads the header section only and writes a portable HTML report for review and sharing.
R package installation
Install the released R package from CRAN:
install.packages("vcfheader")
library(vcfheader)
For more information see the package page, reference manual, and vignettes on CRAN.
Quick start in R
hdr <- parse_vcf_header("file.vcf.gz")
vcfheader(hdr, file = "file_vcfheader.html")
This works with both .vcf and .vcf.gz files.
You can also let vcfheader() derive the output path from the original input file path:
hdr <- parse_vcf_header("file.vcf.gz")
vcfheader(hdr)
The package ships with small example files for offline use, including simple.vcf, sv44.vcf, and a prebuilt example report.
simple_vcf <- system.file("extdata", "simple.vcf", package = "vcfheader")
hdr <- parse_vcf_header(simple_vcf)
vcfheader(
hdr,
file = "simple_vcfheader.html"
)
What the report shows
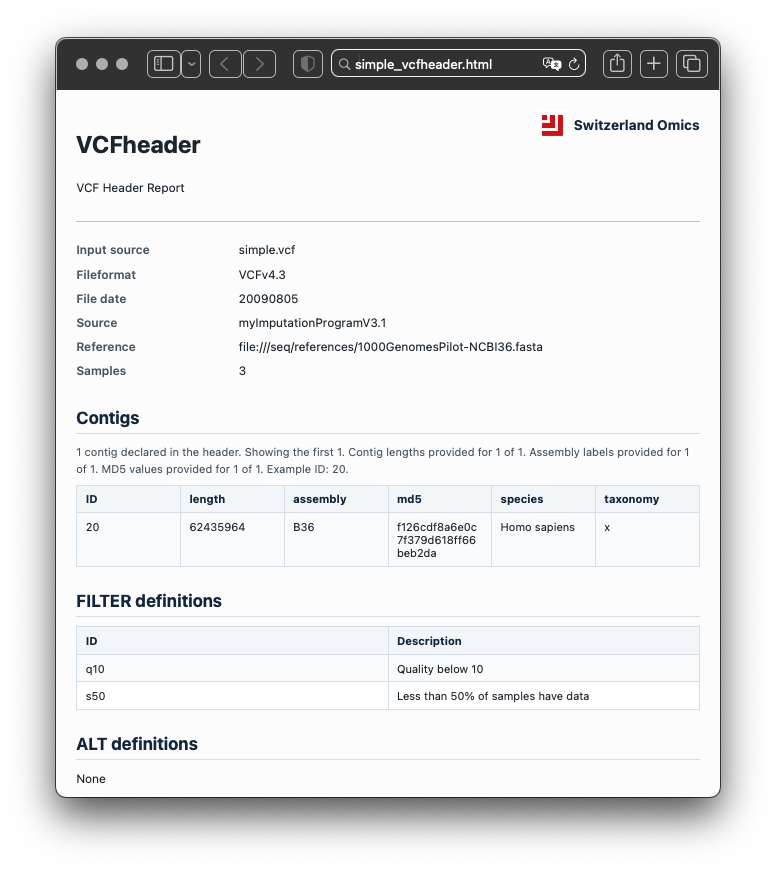
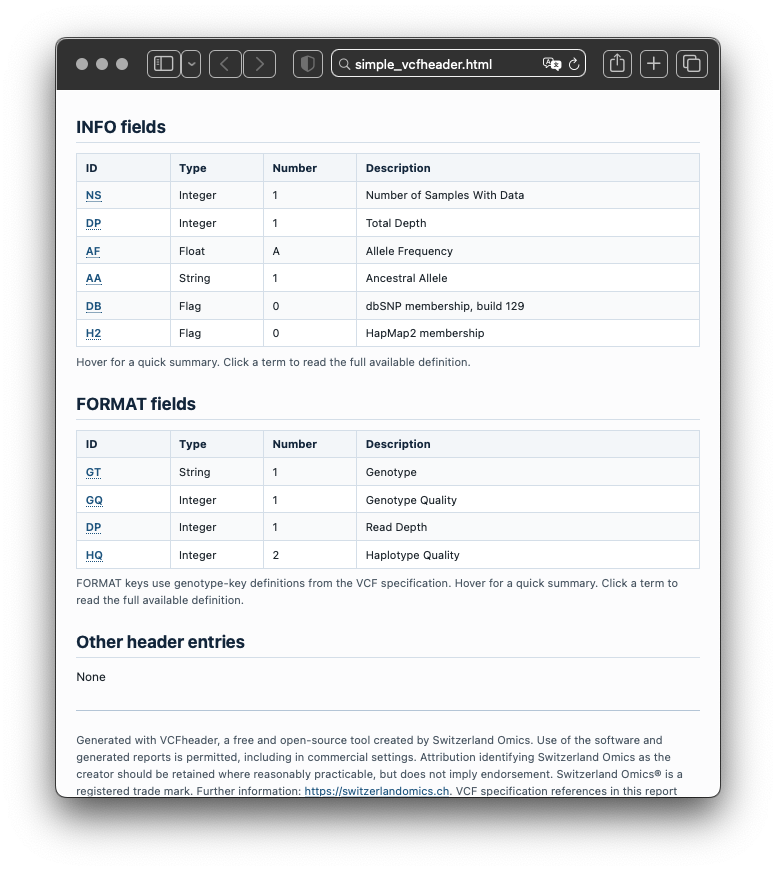
The report summarises file metadata, contigs, INFO fields, FORMAT fields, FILTER definitions, ALT definitions, sample names, warnings, errors, and lightweight inferred annotations. It turns header metadata into a structured record that can be inspected, validated, and shared.
Because only the header is read, review is fast and does not depend on loading full variant records. This makes VCFheader useful for routine checks at the point where a file first enters analysis or review.
Citation
If you use VCFheader, please cite the software release and package source.
R package
Lawless D. vcfheader: Structured parsing and HTML reporting for VCF headers. Comprehensive R Archive Network.
@Manual{vcfheader_cran,
title = {vcfheader: Fast Genetic Variant Call Format File Header Intelligence and Audit},
author = {Lawless, Dylan},
year = {2026},
note = {R package},
url = {https://CRAN.R-project.org/package=vcfheader}
}
Licence
Generated reports are based on VCFheader, a free and open-source tool created by Switzerland Omics. Use of the software and generated reports is permitted, including in commercial settings, under GPL-3. VCF specification references in generated reports may relate to samtools and the broader HTS specifications ecosystem, distributed under the MIT/Expat Licence by Genome Research Ltd. Source: https://github.com/samtools/samtools. Further reading: https://www.htslib.org/doc/#file-formats.


